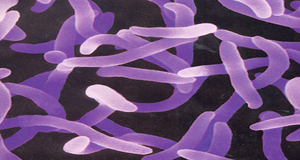

Yersinia pestis, is a nonmotile, nonsporulated, aerobic Gram-negative bacillus or coccobacillus exhibiting a hairpin morphology after Gram staining and growing within 24 to 72 h at a temperature range of 4 to 40°C (optimum, 28 to 30°C) at pH 7.4[1]. Transmitted by fleas from rodent reservoirs, Y. pestis infection may cause five principal forms of plague, including bubonic, septicemic, pneumonic, meningeal, and pharyngeal plague[2]. In addition, Y. pestis may cause skin ulceration at its portal of entry, reported as carbuncles and ulcers, along with pustules, spots, petechiae, bruising, and gangrene[3].
Three historic plague pandemics have caused over 160 million deaths worldwide[1]. The first pandemic, known as the Justinian Plague, devastated the Mediterranean Basin from 541 to 750/767 CE[4] and invaded northern Europe as far as Germany and England[2]. The second pandemic, lasting from 1346 to the 18th century, including the so-called “Black Death” period of 1346 to 1353[2], killed an estimated one-third of the European population[5]. The third pandemic probably began at the end of the 19th century in Hong Kong, China, and then spread to Africa, America, Oceania, and other parts of the world via maritime trade, and lasted until the mid-20th century[6]. At present, plague remains a threat in many parts of the world and has been categorized by the World Health Organization since 2000 as a reemerging disease.
Serotyping and phage typing are not generally useful for differentiating among Y. pestis strains due to a high level of homogeneity[7]. Y. pestis strains have different abilities to ferment glycerol and arabinose and reduce nitrate. Based on this, they have been classified into five biovars (antiqua, mediaevalis, orientalis, microtus and Intermedium)[8]. The strains of the former three biovars are highly virulent to animals or humans, while the biovar microtus strain, including ‘pestoides’ one, is avirulent or opportunistic to larger mammals but virulent to small rodents[9]. Currently, 33 phylogroups in five main branches of Y. pestis are identified. Branch 0 is a root lineage of Y. pestis, which contains several ‘untypical’ groups. A ‘Big Bang’ node of Y. pestis emerged between 1330–1340, with Branch 1–4, radiated from it[10]. Branch 1 is currently most widely distributed lineage of Y. pestis that currently thrives in natural plague foci in Asia, Africa, and America, and most likely also thrives in Europe in the late-medieval and possibly early modern periods[11]. Branch 2 is split into 2.ANT and 2.MED lineages, with the strains of the former circulating in Nepal, China, and Mongolia, while 2.MED is found all over Asia, all the way from Caucasus, Caspian, Volga-Ural region in the west, via western Kazakhstan, Turkmenistan, northern Kyrgyzstan, into China and Mongolia. Strains of Branch 3 were only found in Gansu Province and Qinghai Province of China and Mongolia and strains of Branch 4 were only found in Russia and Mongolia[12].
Y. pestis contains the plasmid pCD1 (70–75 kb)[7] and horizontally acquired two additional plasmids during evolution, pMT1 (100–110 kb) and pPCP1 (9.5kb), and a high pathogenicity island consisting of 32 chromosomal genes that is unique to Y. pestis. Some determinants encoded by plasmids pMT1 and pPCP1 facilitate Y. pestis-specific tissue invasion, survival in flea vectors or possibly its heavy growth in host blood.
The expression of putative virulence locus in Y. pestis (hms, caf1, T3SS and psa) is responsive to a wide range of environmental stresses and multiple regulatory proteins. Other genes responsible for cellular metabolism were also actived upon exposure to multiple stresses, including energy metabolism, sulfur metabolism, ribosome protein biosynthesis, iron uptake, heme synthesis and utilization to chemotaxis and motility[13]. The attachment invasion locus is a chromosomally encoded small-membrane protein and is the primary adhesin of Y. pestis. Yersiniabactin has high affinity for ferric iron and is necessary for acquiring iron from transferrin and lactoferrin by yersiniae[14].
Samples collection date:
Samples host information:
Samples phylogroup information:
Samples Virulence information:
Samples Resistance information:
[1] Perry R D, Fetherston J D. Yersinia pestis - Etiologic agent of plague[J]. Clinical Microbiology Reviews, 1997, 10(1): 35-66.
[2] Lei, Liu, Shangen, et al. Transcriptional regulation of Yersinia pestis biofilm formation[J]. Microbial pathogenesis, 2019, 131: 212-217.
[3] Thompson K M. Environmental Regulation of Yersinia Pathophysiology[J]. Front Cell Infect Microbiol, 2016, 6(25): 25.
[4] Darby C. Uniquely insidious: Yersinia pestis biofilms[J]. Trends in Microbiology, 2008, 16(4): 158-164.
[5] Nuri R, Shprung T, Shai Y. Defensive Remodeling: How bacterial surface properties and biofilm formation promote resistance to antimicrobial peptides[J]. BBA - Biomembranes, 2015: 3089-3100.
[6] Spyrou M A, Bos K I, Herbig A, et al. Ancient pathogen genomics as an emerging tool for infectious disease research[J]. Nat Rev Genet, 2019, 20(6): 323-340.
[7] Barbieri R, Signoli M, Chevé D, et al. Yersinia pestis: the Natural History of Plague[J]. Clin Microbiol Rev, 2020, 34(1): e00044-19.
[8] Qi Z, Cui Y, Zhang Q, et al. Taxonomy of Yersinia pestis[J]. Adv Exp Med Biol, 2016, 918: 35-78.
[9] Zhou D, Han Y, Yang R. Molecular and physiological insights into plague transmission, virulence and etiology[J]. Microbes Infect, 2006, 8(1): 273-84.
[10] Bramanti B, Wu Y, Yang R, et al. Assessing the origins of the European Plagues following the Black Death: A synthesis of genomic, historical, and ecological information[J]. Proc Natl Acad Sci U S A, 2021, 118(36): e2101940118.
[11] Spyrou M A, Tukhbatova R I, Feldman M, et al. Historical Y. pestis Genomes Reveal the European Black Death as the Source of Ancient and Modern Plague Pandemics[J]. Cell Host Microbe, 2016, 19(6): 874-81.
[12] Kutyrev V V, Eroshenko G A, Motin V L, et al. Phylogeny and Classification of Yersinia pestis Through the Lens of Strains From the Plague Foci of Commonwealth of Independent States[J]. Front Microbiol, 2018, 9: 1106.
[13] Han Y, Qiu J, Guo Z, et al. Comparative transcriptomics in Yersinia pestis: a global view of environmental modulation of gene expression[J]. BMC Microbiol, 2007, 7: 96.
[14] Yang R, Atkinson S, Chen Z, et al. Yersinia pestis and Plague: some knowns and unknowns[J]. Zoonoses (Burlingt), 2023, 3(1): 5.