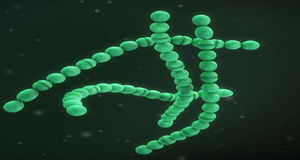

Streptococcus suis (S. suis) is a facultatively anaerobic Gram-positive ovoid or coccal bacterium surrounded by a polysaccharide capsule[1]. This bacterium is a zoonotic agent that causes sepsis and meningitis in pigs and humans. Human infections occur primarily through close contact with infected pigs or through the consumption of raw or undercooked contaminated pork products[1]. This bacterium was first reported by veterinarians in 1954, after outbreaks of meningitis, septicemia, and purulent arthritis occurred among piglets. Fourteen years later, the first human S. suis cases were diagnosed in Denmark[2].
About 1,000 different sequence types (STs) were identified in S. suis based on multi-locus sequence typing (MLST). Worldwide ST1, ST25, and ST28 are the most prevalent STs isolated from swine, the predominance of each ST tends to vary with geographic location[3]. MLST has revealed the presence of many clonal complexes (CCs) within the S. suis population. The most important CCs causing infections in pigs and humans are CC1, CC13/149, CC16, CC17, CC20, CC25, CC28, CC94, CC104, CC233, CC221/234, CC1109, CC1112, and CC1237[4,5]. In North America, CC25 (Canada) and CC28 (United States and Canada) are more commonly reported[6]. These latter CCs are also present in Australia and some parts of Asia[7], whereas CC1 strains are more prevalent in Europe, Asia, and South America[8]. CC20 is important in The Netherlands[9], whereas CC104 and CC233 (ST233, ST379, and ST1656) have caused outbreaks and are endemic to Thailand[10]. CC16 and CC94 predominate among swine isolates in Europe; however, human cases caused by isolates of the latter CCs have also been reported in Thailand[11].
Serotype identification is considered the typing gold standard for S. suis strains. The capsular polysaccharide (CPS) is the basis for the typing system of S. suis[3]. S. suis had originally been classified into 35 serotypes (1/2 and 1–34) based on the antigenicity of the capsular polysaccharide, which is suggested to be a major virulent factor. Because some described serotypes (serotypes 20, 22, 26, 32, 33, and 34) belong to other bacterial species, the number of official S. suis serotypes has been reduced to 29[12]. However, after 2010, serotypes 21/29, NCL21-NCL26 (novel cps loci), and Chz were described in China[13]. Among these, serotype, serotype 2 is regarded as the most virulent and the most frequently isolated serotype infecting humans, causing serious illness and streptococcal toxic shock syndrome (STSS), which was the main cause of death in the 2005 outbreak in Sichuan province of China[14]. STSS-causing S. suis has evolved to acquire, most likely through horizontal gene transfer, an 89 K pathogenicity island (89 K PaI) with multiple virulence genes. The most commonly described virulence markers for S. suis also include muramidase-released protein (MRP), extracellular protein factor (EF), and the hemolysin suilysin (SLY), which were mainly associated with a virulence potential of S. suis serotype 2 strains[5].
Genes encoding resistance against tetracycline, macrolides, aminoglycosides, chloramphenicol, and other antimicrobial drugs have been identified in the various sequenced S. suis genomes. As in other streptococci, many of the resistance genes identified in S. suis have been found to be carried by integrative and conjugative elements (ICEs), transposons, genomic islands, phages, and chimeric elements[8]. Resistance to macrolides, lincosamides, tetracyclines, and sulphonamides has been reported, with up to 85% of strains resistant. S. suis isolates are uniformly sensitive to penicillin and ampicillin[15]. Since 2010, the number of reported S. suis infections in humans has increased substantially; most cases have originated in Southeast Asia, where the density of pigs is high[16] .
Samples collection date:
Samples host information:
Serotype information:
Samples MLST information:
Samples Virulence information:
Samples Resistance information:
[1] Brizuela J, Kajeekul R, Roodsant TJ, et al. Streptococcus suis outbreak caused by an emerging zoonotic strain with acquired multi-drug resistance in Thailand. Microb Genom. 2023;9(2):mgen000952.
[2] Wertheim HF, Nghia HD, Taylor W, Schultsz C. Streptococcus suis: an emerging human pathogen. Clin Infect Dis. 2009;48(5):617-625.
[3] Nicholson TL, Kalalah AA, Eppinger M. Population structure and genetic diversity of Streptococcus suis isolates obtained from the United States. Front Microbiol. 2023;14:1250265.
[4] Hatrongjit R, Fittipaldi N, Gottschalk M, Kerdsin A. Tools for Molecular Epidemiology of Streptococcus suis. Pathogens. 2020;9(2):81.
[5] Scherrer S, Rosato G, Spoerry Serrano N, et al. Population structure, genetic diversity and pathotypes of Streptococcus suis isolated during the last 13 years from diseased pigs in Switzerland. Vet Res. 2020;51(1):85.
[6] Fittipaldi N, Xu J, Lacouture S, et al. Lineage and virulence of Streptococcus suis serotype 2 isolates from North America. Emerg Infect Dis. 2011;17(12):2239-2244.
[7] Segura M, Aragon V, Brockmeier SL, et al. Update on Streptococcus suis Research and Prevention in the Era of Antimicrobial Restriction: 4th International Workshop on S. suis. Pathogens. 2020;9(5):374.
[8] Athey TB, Teatero S, Takamatsu D, et al. Population Structure and Antimicrobial Resistance Profiles of Streptococcus suis Serotype 2 Sequence Type 25 Strains. PLoS One. 2016;11(3):e0150908.
[9] Schultsz C, Jansen E, Keijzers W, et al. Differences in the population structure of invasive Streptococcus suis strains isolated from pigs and from humans in The Netherlands. PLoS One. 2012;7(5):e33854.
[10] Brizuela J, Kajeekul R, Roodsant TJ, et al. Streptococcus suis outbreak caused by an emerging zoonotic strain with acquired multi-drug resistance in Thailand. Microb Genom. 2023;9(2):mgen000952.
[11] Kerdsin A, Hatrongjit R, Gottschalk M, et al. Emergence of Streptococcus suis serotype 9 infection in humans. J Microbiol Immunol Infect. 2017;50(4):545-546.
[12] Obradovic MR, Segura M, Segalés J, Gottschalk M. Review of the speculative role of co-infections in Streptococcus suis-associated diseases in pigs. Vet Res. 2021;52(1):49.
[13] Wang X, Sun J, Bian C, et al. The population structure, antimicrobial resistance, and pathogenicity of Streptococcus suis cps31. Vet Microbiol. 2021;259:109149.
[14] Tan C, Zhang A, Chen H, Zhou R. Recent Proceedings on Prevalence and Pathogenesis of Streptococcus suis. Curr Issues Mol Biol. 2019;32:473-520.
[15] Varela NP, Gadbois P, Thibault C, Gottschalk M, Dick P, Wilson J. Antimicrobial resistance and prudent drug use for Streptococcus suis. Anim Health Res Rev. 2013;14(1):68-77.
[16] Susilawathi NM, Tarini NMA, Fatmawati NND, et al. Streptococcus suis-Associated Meningitis, Bali, Indonesia, 2014-2017. Emerg Infect Dis. 2019;25(12):2235-2242.